
Ajude-me a Vencer
Vote no Seu FinalistaCezar Kayzuka
FMRP-USP - Ribeirao Preto Medical School - University of Sao Paulo
Why should you vote for me!

Abstract
Vascular inflammation can lead to endothelial-to-mesenchymal transition (EndMT), a process whereby endothelial cells lose their normal function and take on mesenchymal phenotypes. This contributes to vascular dysfunction and diseases like hypertension. EndMT involves extensive chromatin remodeling and significant changes in gene expression, linked to the loss of heterochromatin-associated histone H3.1 due to oxidation from inflammation-produced reactive oxygen species (ROS).vWe performed experiments using HULEC-5a (Human Lung Endothelial Cell) treated with TNF-α (10ng/ml), a pro-inflammatory cytokine that induces endMT. Cell lysates were used to measure proteins and mRNA levels by Western Blot and qPCR, respectively. Levels of nuclear ROS were measured using a genetically encoded biosensor, NLS-Orp1roGFP2, stably transfected into HULEC-5a cells. To investigate whether nitric oxide (NO) or derived oxidants were involved in H3.1 degradation, nitric oxide synthase Inhibitors L-NIO (20 μM) and 1400w (70 nM) were used. Whole transcriptome analysis (RNAseq) of cells stimulated with TNFα in the presence and in the absence of NOS inhibitors was performed to detect differential gene expression regulation by TNFα, NO, or both.
In this study, we tested the association between H3.1 loss and EndMT. EndMT was induced in Hulec-5a endothelial cells by treating them with TNF-α for up to 72 hours. The induction of EndMT led to the downregulation of endothelial-specific markers, such as VE-Cadherin and eNOS, and an increase in α -SMA (alpha-smooth muscle actin) and N-cadherin, which are mesenchymal lineage markers. At these same time points, we observed significant increases in nuclear ROS levels, accompanied by the loss of Histone 3.1. The loss of H3.1 was prevented by NOS inhibitors, such as L-NIO and 1400W, indicating a role for NO in this process. Whole transcriptome analysis revealed differentially regulated genes by TNFα, NO, or both, highlighting the importance of NO in chromatin remodeling and gene expression associated with EndMT, once endMT marks (e.g. Collagen type 5, SMAD6, ICAM, ITGB, etc) were identified. Our research suggests that nuclear ROS are involved in chromatin remodeling associated with inflammation-driven EndMT. Targeting NOS to prevent loss of chromatin-associated H3.1 suppresses EndMT in vitro. Further studies are needed to determine if NO-driven H3.1 loss directly regulates genes involved in endothelial marker loss and mesenchymal phenotype gain. This could potentially open new pathways to therapeutically target EndMT in vascular inflammation and prevent cardiovascular diseases like hypertension.
As a nurse, my initial exposure to research during my undergraduate studies centered around nursing education, particularly active learning methodologies. This experience led me to Japan, where I had the opportunity to further explore this research topic. While there, I met a nurse working at an oncology research center, who introduced and inspired me to shift my research focus to the field of pharmacology during my final year of undergraduate studies.
Akira Endo. Unfortunately, he has already passed away (2024), but I admire him for his discovery of the statin that saves countless lives in the cardiovascular context around the world. He also caught my attention due to the coincidence that he was born in the same province as my grandfather, in Akita, Japan.
I can say that I have two personalities since I was born and raised in Japan until 12 years old and then I moved to Brazil. Depending on the day, I wake up with my Japanese personality, when I listen to J-POPs like Arashi and Radwimps and want to eat Japanese foods like Takoyaki and Japanese Curry. When I'm in my Brazilian personality, I enjoy watching soap operas and drinking caipirinha.
Enthusiastic, resilient and curious.
Beatriz Oliveira de Faria
IOC/Fiocruz - Oswaldo Cruz Institute - Oswaldo Cruz Foundation
Why should you vote for me!

Abstract
Slaughterhouse Wastewater Treatment Plants (SWWTPs) are identified as hotspots for the dissemination of antimicrobial resistance, a global health concern. This study presents a novel investigation assessing the effects of advanced oxidation treatment on the removal of both antibiotic-resistant bacteria (ARBs) and antibiotic resistance genes (ARGs) from poultry slaughterhouse effluents, aiming to mitigate the environmental spread of antimicrobial resistance. Samples were collected from the influent and effluent of a poultry SWWTP located in Rio de Janeiro, Brazil.
Effluent samples underwent treatment with UV/H₂O₂ in a benchtop reactor at varied oxidizing agent concentrations and different pH levels. DNA and plasmid DNA (pDNA) were extracted from both influent and treated effluent samples and purified using the Wizard® DNA Clean-Up System (Promega) for qPCR analysis and shotgun sequencing on the MiSeq Illumina platform. Slaughterhouse wastewater harbors plasmid-linked ARGs associated with clinically relevant bacterial orders, such as Enterobacteriales. The SWWTP exhibited inefficient removal of these elements, emphasizing the role of slaughterhouse effluents in environmental AMR dissemination. UV/H₂O₂ treatment at pH 3 and 7 was notably effective in removing plasmids and reducing ARGs in the effluent. These results demonstrate that UV/H₂O₂ treatment is a potent strategy for reducing pathogen-associated ARGs.
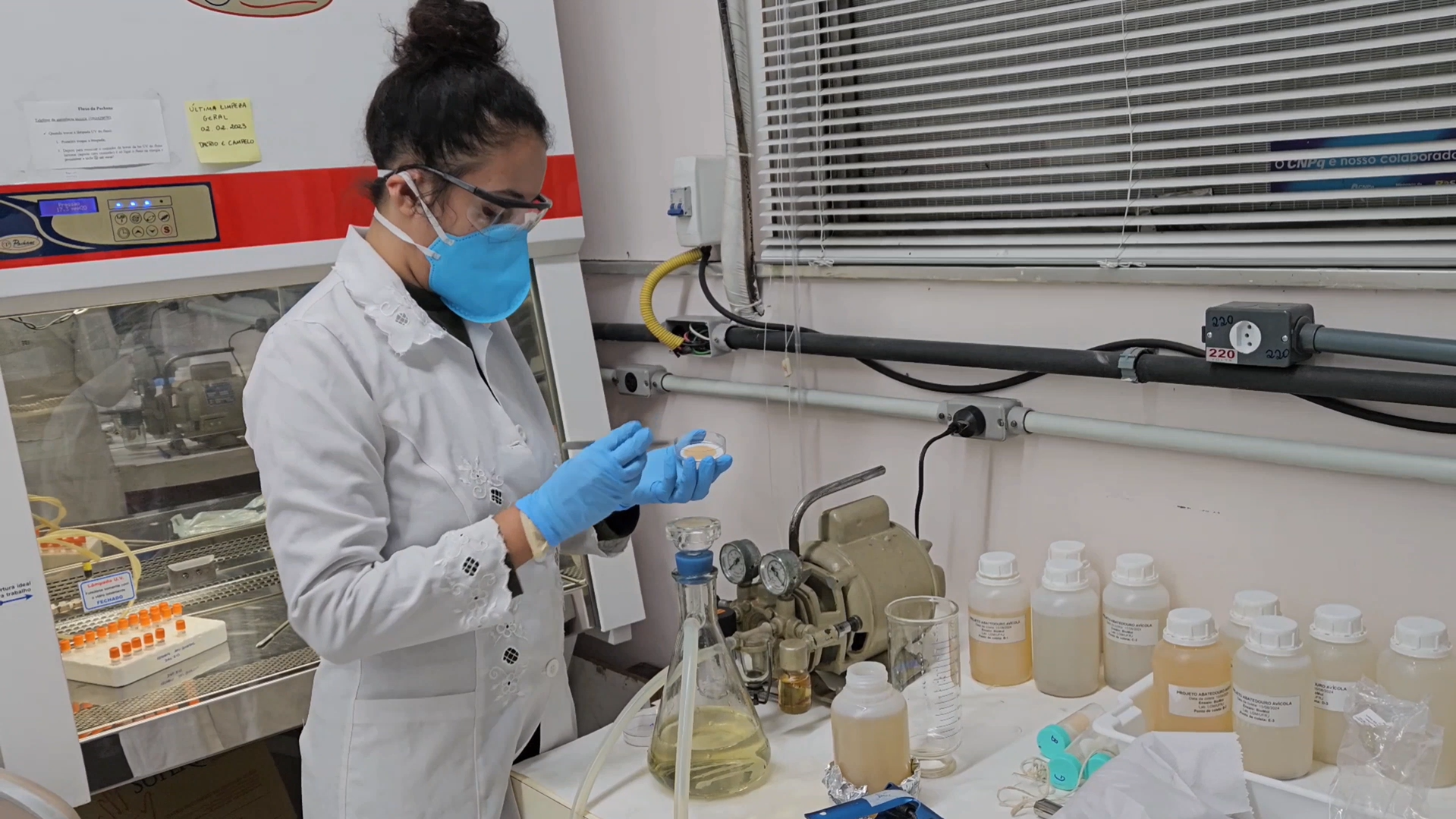

My scientific journey has roots in my childhood, spent on an island surrounded by lush nature and rich history, where my family is part of the local traditional community. I grew up exploring the flora and fauna around me, which sparked my interest in environmental preservation. Although I had no contact with scientists as a child, I always said I wanted to become one because I was fascinated by the natural phenomena I observed around me. This unique experience shaped my perspective and commitment to science and sustainability.
A fun fact about me is that I was literally born on a boat! I've always had a close connection with the sea and developed a deep passion for sailing, especially sailboats. This bond with the ocean has taught me to face challenges with patience and determination. When I’m not working on scientific projects, you’ll most likely find me sailing, enjoying the peace and freedom that only the sea can offer.
Brave, determined and authentic.
Alexandre Luiz Korte de Azevedo
UFPR - Federal University of Paraná
Why should you vote for me!

Abstract
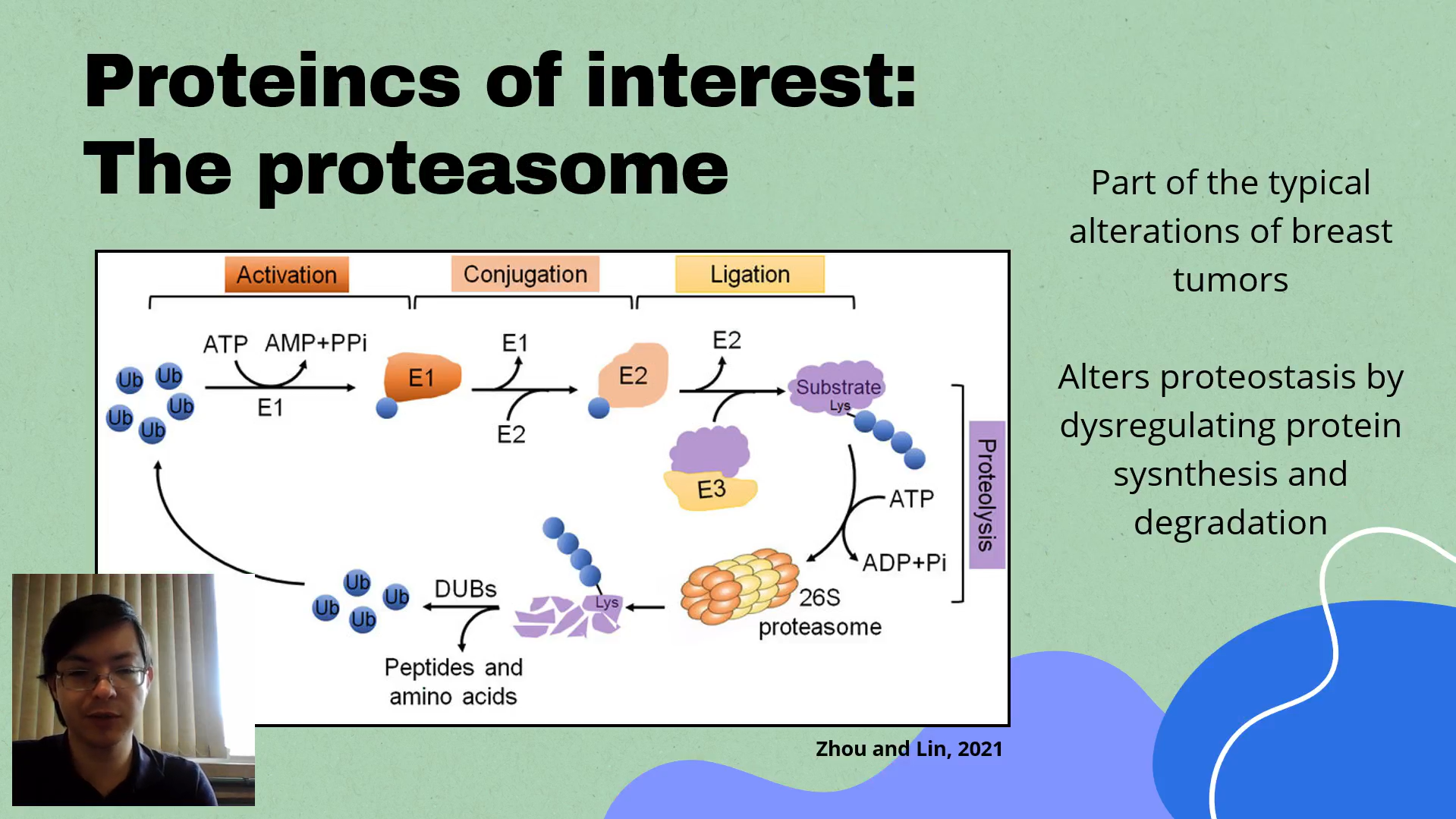
Breast cancer is the most diagnosed and lethal cancer type among women worldwide, yet, despite recent advances in clinic research, therapies that present good efficacy and low toxicity and side effects are still needed to improve patient survival and overall life quality. The chaperone and proteasome networks, both part of the proteostasis system, are dysregulated in breast cancer and contribute to its malignancy and aggressiveness. In fact, proteasome inhibitors, such as oprozomib and bortezomib, have presented high potential as therapeutic options for breast cancer impairment. However, due recurrent drug-resistance, its efficacy is reduced. It this project, in which I participate as the main responsible for the experimental procedures, our objective is to explore the proteomic and transcriptomic landscape of the chaperone and proteasome networks in the disease and investigate the mechanisms responsible for the dysregulation of both systems in breast cancer, aiming to contribute to better understand its impact in breast cancer clinical and biological behavior, besides possible mechanism of proteasome-inhibitors resistance. Initially, we used bioinformatic approaches to predict transcriptional regulators of the proteostasis genes in breast cancer, and found NFY-A as a major regulator of chaperones, co-chaperones, and proteasome subunits.
We validated it impact in proteostasis gene expression by qPCR, and evaluated its impact in cell phenotypes by silencing NFY-A in vitro by siRNA and evaluating cell viability, proliferation, and colony formation, obtaining significant results (P-value < 0.05). After, we selected STIP1, a co-chaperone regulated by NFY-A, and performed a co-immunoprecipitation associated to mass-spectrometry to evaluate its interactome. We identified among its interaction partners the proteasome subunits PSMB2, PSME2, PSMB1, and PSMD2, and chaperones such as HSP90AB1, HSPA5, and HSP90B1. With our partial results, we could identify a common transcription regulator associated to the dysregulation of both chaperone and proteasome networks in breast cancer, shedding light on the origin of the super-activation of both systems in the disease. Also, we demonstrate that the proteostasis genes regulated by NFY-A compose an interactome, indicating a possible mechanism of interdependence of both systems. In the next steps, we intent to evaluate the proteasome activity in NFY-A silenced cells using the cell-based Proteasome-Glo™ assays (PROMEGA G8760), and determine the relationship between the aforementioned chaperones/co-chaperone with the activity and stability of the proteasome in vitro. In general, our project has potential to contribute to the understanding why breast tumors present dysregulated proteostasis and unravel the interplay of the chaperone and proteasome networks in the disease, providing insights of possible mechanisms of proteasome-inhibitors resistance.
While there are many renowned scientists whose contributions to the society are undeniable, like Carlos Chagas or Rosaling Franklin, my favorite scientist is my former advisor, Dr. Iglenir João Cavalli. His mentorship and commitment to community have not only shaped my academic journey but have also inspired me to become a better scientist. The impact of his contributions to our university research and into mentoring new researches, while always being an approachable and pleasant person, is encouraging. I am truly grateful for his influence in my life.
I really thought that my trajectory in my current laboratory would end before it began. Why? Because in the form that I was supposed to submit to be able to participate in the selection of new interns in the laboratory, I misspelled the name of one of the recruiting supervisors. It was a strange mistake, which completely changed the pronunciation of the name. In the weeks before the selection I thought that name was correct. I only discovered the unfortunate error on the day of the interview, after realizing that the name on the office door was different from what I had written on the form. I almost gave up on the interview because I was so embarrassed. How could I have misspelled the name of one of my potential supervisors? But I'm glad I didn't give up! I passed the interview, completed the internship, master's degree, and now I'm in my doctorate!
Open-minded, resourceful and curious
Lídia do Nascimento Cavalcante
UNICAMP - State University of Campinas
Why should you vote for me!

Abstract
Citrus canker disease is caused by Xanthomonas citri subsp. citri (Xcc), a bacterial pathogen that relies on the translocation of effector proteins called Transcription Activator-Like Effectors (TALEs) into host cells to successfully colonize citrus plants. The Xcc TALE PthA4, which is essential to cause canker, directly transactivates CsLOB1, a major citrus canker susceptibility gene. Recent studies have revealed that CsLOB1-INDUCED EXPANSIN 1 gene (CsLIEXP1) is highly and directly upregulated by CsLOB1 in Xcc-infected plants. Because expansins are associated with cell wall loosening, potentially facilitating bacterial colonization, the CsLOB1-dependent activation of CsLIEXP1 is thought to contribute to canker symptoms and disease progression. Thus, CsLIEXP1 likely represents a critical canker susceptibility gene. In this study, we employed CRISPR/Cas9 to disrupt the function of CsLIEXP1 by modifying its corresponding coding region in Citrus sinensis cv ‘Hamlin’ and evaluated the post-infection responses of edited plants.
The Agrobacterium-mediated transformation of epicotyls was performed using the EHA105 strain harboring the pDIRECT_22c vector carrying three sgRNAs targeting the first CsLIEXP1 exon. Sanger sequencing confirmed that CsLIEXP1-edited plants exhibited 26.4% of edited sequences. Three types of indels were obtained: a deletion of a thymine, a deletion of five nucleotides, and the insertion of a thymine CsLIEXP1-edited plants. For bacterial infection assays, leaves of edited and control plants were infiltrated with Xcc-GFP cell suspension at 104 UFC.mL-1 and analyzed by fluorescence and light microscopy 14 days after inoculation. Leaves of edited plants showed less canker symptoms with lesions limited to the site of bacterial inoculation and less pronounced cellular hypertrophy compared to control plants. Furthermore, CsLIEXP1 protein accumulation was reduced in CsLIEXP1-edited plants compared to wild-type plants when infected with Xcc.
These findings strengthen the notion that reduction of CsLIEXP1 protein levels confers tolerance to Xcc infection possibly by a mechanism involving reduced cell growth and expansion. Considering that CsLOB1 is both a susceptibility gene and a transcription factor associated with growth and development of plant organs, silencing of CsLOB1can be detrimental to plant growth, with potential impacts on development and productivity. Novel canker susceptibility genes downstream of CsLOB1, such as CsLIEXP1, can be better CRISPR targets for citrus canker control. Our results highlight the role of CsLIEXP1 as a canker susceptibility gene and provide new insights into plant-pathogen interactions.

Ever since I was young, I have always received strong encouragement to study and chase my goals. I followed my dreams with determination and overcoming the challenges of leaving Acre state to live in Minas Gerais state for my master’s studies at Federal Universiry of Viçosa and later in Campinas city for my Ph.D., were I lived significant and challenging experiences. My Ph.D. project, focused on genome editing, led to two awards at different scientific conferences. This has been a great satisfaction, as it shows that my work is making a positive impact on the scientific community. This journey is proof that, with dedication and passion, we can reach places that once seemed unimaginable.
Jennifer Doudna and Emmanuelle Charpentier developed the CRISPR-Cas9 technology, which revolutionized genetic engineering and genome editing. They were awarded the Nobel Prize in Chemistry in 2020, becoming two of the 57 women to receive a Nobel Prize, in contrast to a total of 873 men up to that point. The careers of Doudna and Charpentier exemplify the importance of collaboration among scientists to foster new discoveries in scientific fields and drive the development of technologies that benefit humanity. Their stories also highlight the importance of having qualified women in academic environments, which are still male-dominated and where female recognition is often insufficient.
I’m a person who loves animals, especially my little dog Simba. I enjoy going to the gym and love dining at good restaurants with friends.
Kimberly Freitas Cardoso
IRR/Fiocruz - René Rachou Institute - Oswaldo Cruz Foundation
Integrated Biomarker Research Group (GIPB) and Immunology of Viral Diseases
Why should you vote for me!
People should vote for me because my work on the development of a bivalent vaccine against influenza and pneumococcal pneumonia offers an innovative approach to tackling two of the leading causes of global mortality. My research combines cutting-edge technologies with a strong commitment to public health, has the potential to prevent millions of victims of this respiratory diseases worldwide and the award would significantly contribute to the progress of my study.

Abstract
S. pneumoniae and the influenza virus are respiratory tract pathogens that cause high mortality worldwide. The influenza virus causes the flu, while the pneumococcus causes pneumonia and meningitis, as well as secondary infections in flu patients, worsening the clinical condition. Licensed pneumococcal vaccines are serotype-specific, which results in the replacement of circulating serotypes by non-vaccine serotypes. Therefore, aiming to develop a bivalent vaccine against these two pathogens, we constructed a recombinant influenza virus carrying the pneumococcal protein PspA (Flu-PspA), by reverse genetics. Thus, the objective of this work was to evaluate the effectiveness of a vaccine protocol using the Flu-PspA virus and recombinant PspA protein, in a murine model, via the intramuscular route. For this, C57BL/6 mice were immunized with Flu-PspA (1st dose) and with PspA protein adjuvanted with Alum (2nd dose), in the vaccine group; control virus Flu-CT (1st dose) and PspA protein adjuvanted with Alum or only Alum (2nd dose), or two doses of PBS, in the control groups. Fourteen days after the 2nd dose, anti-PspA and anti-influenza IgG antibodies were quantified by ELISA and the ability of anti-PspA antibodies to deposit complement (C3) on the surface of different pneumococcal strains was evaluated by flow cytometry.
Furthermore, 21 days after the last dose, the animals were lethally challenged with 7xLD50 of the invasive pneumococcal strain ATCC6303 or 100xLD50 of the influenza A/PR8/34 virus and survival monitored for ten days. Furthermore, three days after the lethal challenge, the bacterial load in bronchoalveolar lavage fluid (BALF) and blood were determined by titration and inflammatory cytokines in BALF were quantified by CBA. Thus, we observed that the animals in the vaccine group presented high levels of anti-PspA and anti-influenza IgG antibodies in the serum and the anti-PspA antibodies were effective in depositing complement in different pneumococcal strains, demonstrating a broad-spectrum vaccine response. Furthermore, we observed 100% protection after lethal challenges with pneumococcus and influenza, a reduction of two to three logs10 in pulmonary bacterial load and lower levels of inflammatory cytokines in BALF. Therefore, this vaccine protocol showed high protection and immunogenicity, being a promising bivalent vaccine strategy against influenza and pneumococcal pneumonia.
The present work deals with an innovative bivalent vaccine capable of protecting against the influenza virus and providing cross-protection against different pneumococcal serotypes. The technology is original and innovative and has a national application for invention to INPI with process number BR 1020220151636. Furthermore, it has a social and technological impact and can benefit the health of the Brazilian population, especially the portion of the population most susceptible and affected by the two diseases: children under 5 years of age, elderly , pregnant women, people with chronic diseases (e.g. asthmatics, people with obstructive lung diseases, heart disease).
A defining moment in my scientific journey occurred in 2020, when my master’s research was presented to the Brazilian Minister of Science and Technology as a successful example for securing funding to develop a bivalent vaccine against influenza and COVID-19 at the onset of the pandemic in the country. During that time, I was part of the team responsible for developing this vaccine and participated in a feature about the project, which aired on the largest free-to-air TV network in Brazil. In the same year, I was recognized as a standout fellow by CAPES, acknowledging the impact of my research. These achievements not only highlighted the importance of my work but also motivated me to continue seeking innovative solutions to public health challenges, resulting in several awards throughout my career.
I greatly admire the late scientist Rosalind Franklin, whose work in capturing images of the DNA double helix went unrecognized. Her career highlights the historical challenges women have faced in science, which inspires me to contribute to changing this scenario.
A fun fact about me is that, since childhood, my family nicknamed me "Kika" after the character from the Brazilian cartoon series "De onde vem?" because I always had an insatiable curiosity about the origin of things around me, like the character Kika from the cartoon. I also have a deep love for science fiction and am a fan of health-related documentaries and shows, such as "My 600-lb Life," which follow individuals on their journey to overcome obesity. These topics spark my interest in areas beyond my scientific specialty. Recently, I discovered gardening as a relaxing hobby that provides a pleasant break from my daily activities.
Curious, determined, and persistent.
Elevando o seu Caminho à Descoberta
O Prêmio Promega Rising Research tem como objetivo capacitar e reconhecer você e suas contribuições científicas e sua jornada acadêmica como estudante de doutorado. Queremos apoiá-lo no seu desenvolvimento profissional e ajudá-lo a ter acesso a novos recursos, novas perspectivas e novas conexões que ampliarão seus horizontes.
Para ver os países participantes e obter mais informações, visite a página inicial do Rising Researchers Award.
O Brasil é um dos países participantes.

Depoimento de um(a) Ganhador(a)
"Durante o meu mestrado, já tinha colaborado virtualmente com o grupo de Target Engagement da Promega, e foi muito emocionante conhecer pessoalmente este grupo, visitar o seu laboratório, conversar e aprender com eles. Esta rica experiência individual com estes cientistas ajudou a reafirmar o meu interesse na minha área e contribuiu para a minha decisão de prosseguir para um doutorado."
Rebeka Fanti concluiu seu Mestrado pela Unicamp, e atualmente é Aluna de Doutorado no Structural Genomics Consortium, University of Toronto, Canadá.


2º Lugar

3º Lugar
Indivíduos elegíveis não são obrigados a comprar produtos da Promega ou pagar qualquer taxa para participar. Funcionários da Promega Corporation e suas subsidiárias e distribuidores autorizados, assim como membros das famílias imediatas desses funcionários, não são elegíveis.
O Prêmio Promega Rising Researchers não está disponível em todas as regiões geográficas. Os candidatos devem verificar o site da Promega ou consultar seu representante da Promega ou distribuidor autorizado da Promega sobre a disponibilidade em sua área. Nulo onde proibido por lei.
O Prêmio Promega Rising Researchers está disponível para indivíduos que têm pelo menos 18 anos e são pesquisadores ou cientistas matriculados em um programa de doutorado em ciências biológicas na data em que sua inscrição para o Prêmio é submetida. Exemplos de áreas de pesquisa em ciências biológicas elegíveis incluem, mas não se limitam a: biologia, biologia molecular, biotecnologia, bioquímica, ciências biomédicas, genética, microbiologia, farmacologia, neurociência, ecologia, imunologia e outras áreas de pesquisa semelhantes..

Conte-nos um pouco sobre você
Nenhuma inscrição será aceita após 30 de junho de 2024. Cada candidato será notificado se foi selecionado como finalista até, no máximo, 31 de julho de 2024. Cada finalista será notificado se foi escolhido como destinatário até, no máximo, 6 de dezembro de 2024.
O seguinte ilustra as informações que o candidato será solicitado a fornecer ao enviar a inscrição para o programa de premiação.
- Informações de Contato: Nome, sobrenome, data de nascimento, endereço de e-mail, telefone, cargo/função, instituto, endereço, cidade, estado/província (se aplicável), código postal/CEP, país..

Compartilhe seu resumo e o impacto do projeto
Por favor, considere que a política interna de algumas organizações pode não permitir que o candidato receba incentivos ou que a permissão do empregador pode precisar ser solicitada antes de participar do concurso.
Por favor, tenha em mente que este resumo será compartilhado publicamente por meio do site da Promega e considere o nível de informação permitido pela sua organização.

Os 5 melhores finalistas de cada filial participante da Promega
(Selecionados pelos Representantes da Promega)
Os finalistas serão selecionados por meio de um processo de triagem envolvendo funcionários da filial da Promega (o “Comitê de Seleção”) com base nas informações enviadas no formulário de inscrição. Cada destinatário será notificado por e-mail a critério exclusivo da Promega, até quarta-feira, 31 de julho de 2024. No caso de um destinatário não poder receber seu prêmio por qualquer motivo, o prêmio será concedido a um vencedor alternativo conforme determinado pelo Comitê de Seleção da Promega.

Os votantes devem ter um endereço de e-mail com domínio acadêmico
(Os finalistas podem promover seus vídeos para coletar votos)
Todos os finalistas selecionados pelo Comitê de Seleção da Promega serão solicitados a enviar informações adicionais, como uma foto de perfil e respostas a perguntas sobre seu histórico científico e interesses. Os finalistas também serão convidados a preparar um vídeo de até 3 minutos detalhando seu projeto.
Essas informações estarão disponíveis no site da Promega, juntamente com um formulário que permitirá que outros pesquisadores acadêmicos votem em um finalista de sua escolha. O único critério para votar será fornecer o próprio endereço de e-mail acadêmico, ou seja, um e-mail com um domínio acadêmico, como @harvard.edu, @stanford.edu, etc. Os votos serão contados de acordo com o país de residência dos votantes, e um finalista com o maior número de votos por cada filial participante da Promega (e os países que elas atendem) será considerado o destinatário daquela filial da Promega. Portanto, o número de destinatários corresponde ao número de filiais participantes da Promega (e não ao número de países participantes).
Cada destinatário será notificado por e-mail a critério exclusivo da Promega, até sexta-feira, 6 de dezembro de 2024.

O finalista com o maior número de votos de cada filial da Promega vence
(Prêmio principal: viagem a Madison, WI, para conhecer nossa P&D)
Quanto aos custos relacionados ao passaporte do ganhador e ao visto para os EUA, consulte sua filial local da Promega.
Despesas adicionais incorridas pela escolha individual de cada ganhador, como atividades e compras fora da programação da visita, não serão cobertas pela Promega.
Premiações adicionais para outros finalistas estão a critério da filial local da Promega desses finalistas.
Ao aceitar o Prêmio Promega Rising Researchers, o candidato será solicitado a compartilhar informações ou dados coletados para fins de marketing ou comerciais. Especificamente, os vencedores concordam em:
- Compartilhar histórias de sua pesquisa com a Promega nos meses seguintes à recepção do prêmio..
- Apoiar atividades promocionais a pedido da Promega durante 12 meses após a aceitação do prêmio, que podem incluir, mas não se limitam a, entrevistas, apresentações em webinars, destaque de histórias no site, vídeos, sessões de fotos, blogs, compartilhamento de dados e/ou apresentação em conferências ou seminários.
Esse conteúdo será compartilhado no site da Promega (www.promega.com), no blog da Promega (www.promegaconnections.com) e nas contas de mídia social da Promega, incluindo Facebook, Instagram, LinkedIn e Twitter.
Os ganhadores devem reconhecer a Promega em publicações científicas de autoria do vencedor relacionadas à pesquisa realizada e apoiada pelo prêmio. Um formulário de consentimento detalhado será fornecido, permitindo que os candidatos escolham quais informações estão dispostos a compartilhar e por quais canais.
Em caso de disputa quanto à identidade da pessoa que enviar uma inscrição online, a inscrição será considerada como tendo sido feita pela pessoa cujo nome está registrado na conta de e-mail. O destinatário pode ser solicitado a fornecer provas de que é o titular autorizado da conta de e-mail associada à inscrição selecionada. O retorno de qualquer notificação de prêmio como não entregue resultará na perda do prêmio.
As informações de inscrição serão propriedade da Promega. Nenhuma transferência de prêmio ou resgate em dinheiro será permitida. Nenhuma substituição de prêmio será permitida, exceto a critério exclusivo da Promega, caso em que um prêmio de valor comparável ou maior poderá ser concedido.
A Promega reserva-se o direito de substituir qualquer prêmio por um de valor igual ou maior ou de cancelar, suspender e/ou modificar o prêmio a seu exclusivo critério.
Ao participar, os candidatos concordam em respeitar e se submeter às regras e decisões da Promega, que serão finais em todos os aspectos relacionados a este Prêmio Promega Rising Researchers, incluindo, sem limitação, a interpretação desta regra.
Os participantes concordam em isentar, liberar e isentar a Promega, suas subsidiárias, afiliadas, diretores, agentes, representantes e respectivos funcionários de qualquer e todas as reclamações, cobranças, lesões, responsabilidades, perdas e/ou danos de qualquer natureza resultantes ou decorrentes da participação no Prêmio Promega Rising Researchers e/ou da aceitação, uso, má utilização ou posse de quaisquer produtos recebidos através do Prêmio Promega Rising Researchers. Os destinatários deste prêmio não serão elegíveis para futuros.